Section: New Results
Classification of diffusion dynamics from particle trajectories
Participants : Vincent Briane, Charles Kervrann.
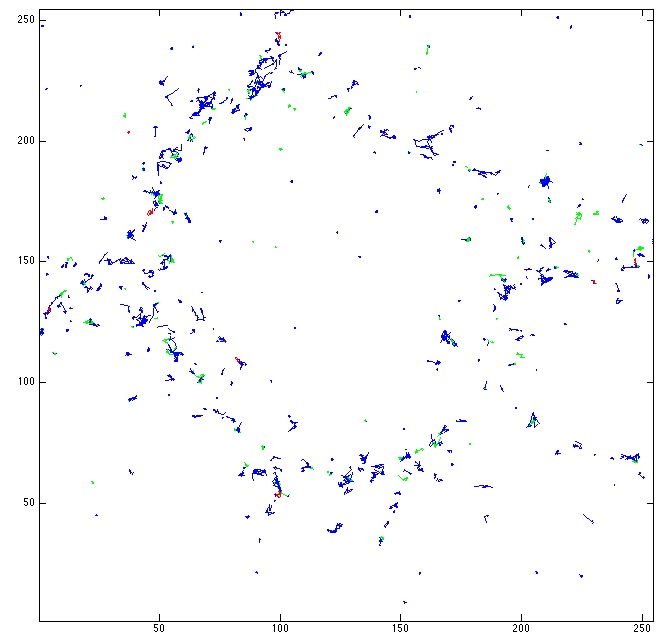
In this study, we are currently interested in describing the dynamics of particles inside live cell. We assume that the motions of particles follow a certain class of random process: the diffusion processes. In 2015, we developed a statistical test to classify the intracellular motions into three groups : free diffusion (i-e Brownian motion), subdiffusion and superdiffusion. This method is an alternative to the commonly used Mean Square Displacement (MSD) analysis. This year, we have studied theoretical properties of our procedure. We have shown that it behaves well asymptotically, that is when we observe the particle trajectory for a very long time, for certain parametric models. The models on which we assess our procedure are representative of the three classes aforementioned and extensively used in the literature. Among them we can cite Brownian motion with drift, Ornstein-Uhlenbeck process and fractional Brownian motion. An illustration of the testing procedure is shown in Fig. 7.
We also extend our method to address two different questions. First, we are interested in testing a large number of trajectories. The first version of our test is a single test procedure. It is known that applying multiple times a test without care leads to a high number of false positives. Then, we modify our initial method to overcome this problem. Secondly, in the case in which we observe very long trajectories, it is likely that the particle motion changes over time. Therefore, we are currently adapting our initial procedure to detect change-point along a single trajectory.
Collaborators: Myriam Vimond (ENSAI Rennes),
Jean Salamero (UMR 144 CNRS-Institut Curie).